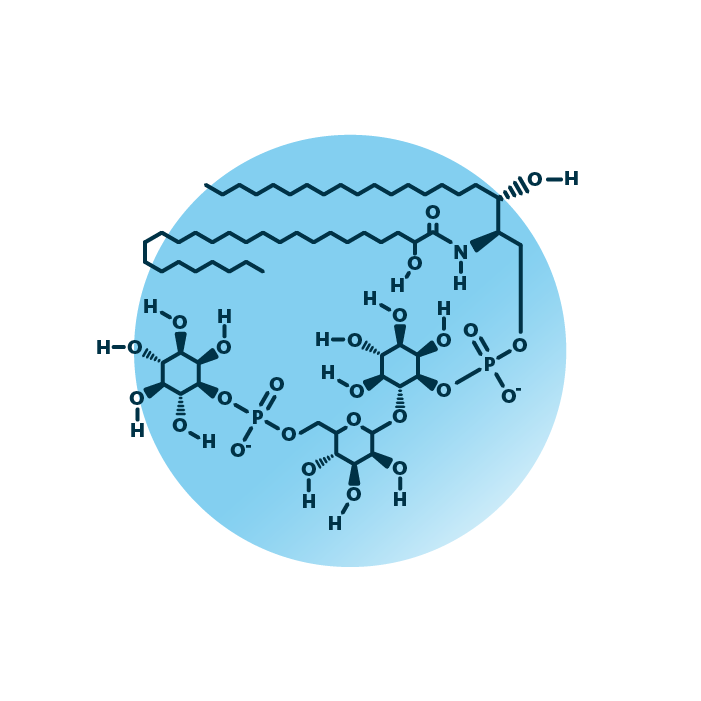
About the structure and biological function of M(IP)2C
Structure. Mannosyl-di-(inositol-phosphoryl)-ceramides (mannose-(inositol-P)2-ceramides, or M(IP)2C) belong to the group of phosphosphingolipids within the sphingolipids. Their structure consists of a ceramide backbone bound to two linked phosphorylinositol molecules of which one is mannosylated. The ceramide backbone contains two hydrocarbon chains: a long-chain base which is linked to a fatty acid via an amide bond. The fatty acid and the long-chain base can be of variable length, hydroxylated, and contain double bonds.
Function. Little is known about the biological function of mannosyl-di-(inositolphosphoryl)-ceramides but they are important components of biological membranes of fungi. Together with MIPC, they constitute the major sphingolipids in yeasts. As M(IP)2C lipids are not produced in mammals, targeting their synthesis is a strategy for novel antifungal drugs. Further, M(IP)2C ceramides interact with plant defensins, small peptides produced by plants that possess antifungal activity.