About the structure and biological function of Gb3
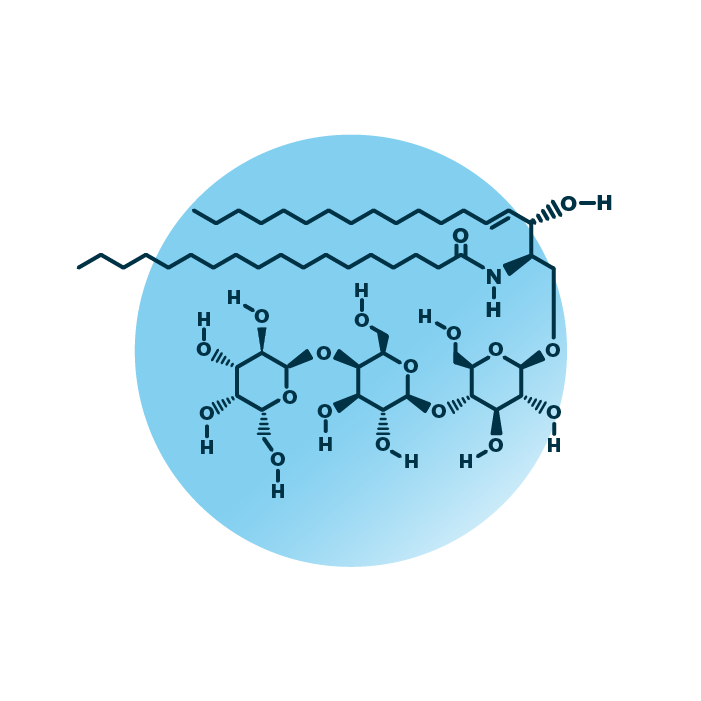
Structure. Gb3 globoside lipids (globo-triaosylceramides, ceramide trihexosides, or antigen CD77) belong to the group of globosides within the sphingolipids. Their structure consists of a ceramide backbone linked to a neutral oligosaccharide unit made of three sugar molecules. The ceramide backbone contains two hydrocarbon chains: a long-chain base which is linked to a fatty acid via an amide bond. The fatty acid and the long-chain base can be of variable length, hydroxylated, and contain double bonds.
Function. Gb3 globosides specifically interact with the cellular “death receptor” FasR. When paired to a Gb3, FasR transduces a cell death signal derived from activation of the caspase-8 cascade. A genetic defect in the enzyme alpha-galactosidase leads to accumulation of Gb3, causing Fabry’s disease, a lysosomal storage disease. Further, Gb3 are a component of the cell membrane where they bind to Shigatoxins facilitating their entry into cells.